AmpliCHECK - Amplicon Sequencing CHECKing tool
AmpliCHECK is intended for a preliminary analysis of amplicon sequencing results.
Before optimizing parameters for accurate genotyping, we should know the length, coverage and frequency
of the most abundant variants in each amplicon and the potential erroneous ones (artefacts derived from
sequencing or PCR errors).
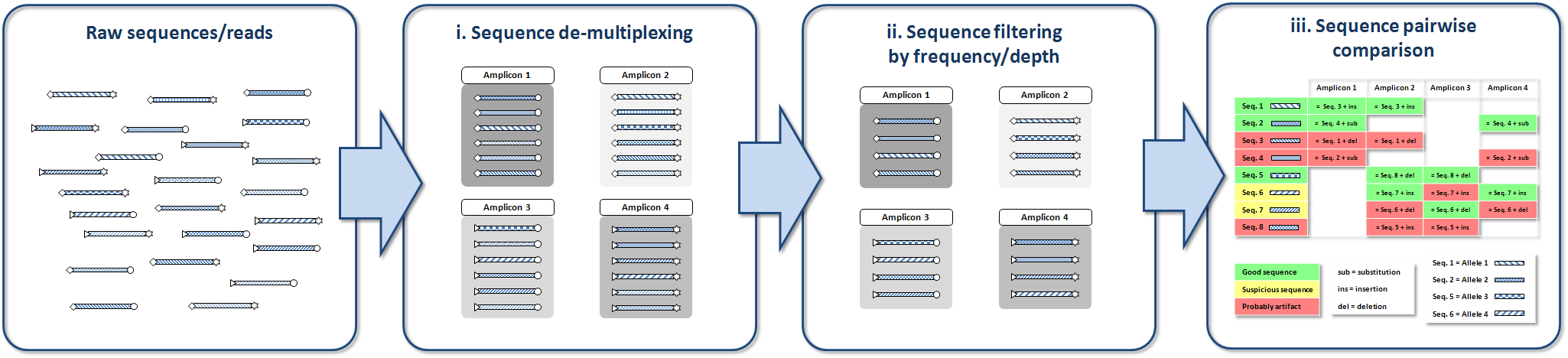
AmpliCHECK de-multiplexes reads and performs a fast analysis of the higher
frequency variants in each individual amplicon retrieving the previously mentioned information.
After running AmpliCHECK we should be able to establish the length of the desired PCR products (markers),
the % of errors in variants and a threshold frequency to decide if a variant is real or is an artefact.
AmpliCHECK takes as input:
- SEQUENCE FILE: FASTQ or FASTA format file (compressed or uncompressed). Multiple sample/amplicon sequence files should be packed into a unique .ZIP or .TAR.GZ file. Also previously analyzed results can be used as input in AmpliSAS format Excel file.
- AMPLICON DATA 1: primer and tag information in a CSV format file as explained in the documentation.
An optional FASTA file with custom allele names and sequences can be provided.
Results can be downloaded on the same page or from an email message after analysis completion.
Results include an Excel file with three spreadsheets, the first includes coverage values, the second, amplicon frequencies and the third, potential sequencing errors and chimeras annotations.
In every spreadsheet, true sequences are colored in green, questionable ones in yellow and clear artefacts in red.
For more information, read the documentation.
Run AmpliCHECK
Disclaimer
Your use of any of these tools is at your own risk. We do not give any representation or warranty nor assume any liability or responsibility for the data nor the results posted (whether as to their accuracy, completeness, quality or otherwise). Access to these data is available free of charge for ordinary use in the course of research. By visiting the site, you accept our use of cookies and you accept that your data and results will be stored in our server.