AmpliCANCER - CANCER variants discovery tool
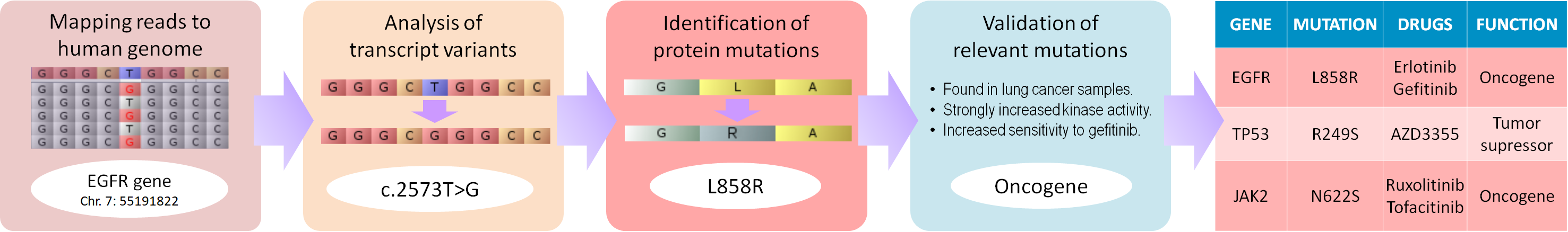
AmpliCANCER analyzes in one-click raw reads from Exome Sequencing, RNA-Seq and Targeted Gene Panels to detect clinically relevant genotypes and reports a full description of the genetic alterations present in the tumor cells together with other important information that could guide in the treatment choice.
Sequencing data is automatically aligned to the latest human reference genome (Ensembl GRCh38.p10).
Genetic variants are scored, filtered, translated and compared against UniProt references, annotating the genetic aberrations detected.
AmpliCANCER has been developed for research purposes and it is not intended for human diagnosis.
AmpliCANCER takes as input:
- SEQUENCE FILE: FASTQ or FASTA format file (compressed or uncompressed). Should contain reads from only one sample/individual.
Results can be visualized on the same page or from an email message after analysis completion.
For more information, read the documentation.
Run AmpliCANCER
Disclaimer
Your use of any of these tools is at your own risk. We do not give any representation or warranty nor assume any liability or responsibility for the data nor the results posted (whether as to their accuracy, completeness, quality or otherwise). Access to these data is available free of charge for ordinary use in the course of research. By visiting the site, you accept our use of cookies and you accept that your data and results will be stored in our server.